AbClon

Antibody engineering services
Antibody Sequencing
Identification of antibody gene sequence information is an essential process for functional opti-
mization, database banking and patent applications antibody engineering, such as humanization and
affinity maturation of monoclonal antibodies (mAbs) . It is a service that can strongly recom-
mend even if cell line management is difficult after Hybridoma development. Hybridoma can
contain pseudogenes and mRNA encoding non-specific antibodies, which can be amplified by
primers to clone the target antibody, after sequencing 3 to 5 clones independent of the
antibody, it is produced in the form of a Fab and reacts with the antigen by ELISA to ensure
accuracy.
mization, database banking and patent applications antibody engineering, such as humanization and
affinity maturation of monoclonal antibodies (mAbs) . It is a service that can strongly recom-
mend even if cell line management is difficult after Hybridoma development. Hybridoma can
contain pseudogenes and mRNA encoding non-specific antibodies, which can be amplified by
primers to clone the target antibody, after sequencing 3 to 5 clones independent of the
antibody, it is produced in the form of a Fab and reacts with the antigen by ELISA to ensure
accuracy.
Features
- Identify variable region sequences that react with antigens by Fab construction
- Duration: about 6 weeks
- Rich experience in mAb sequencing
Customer Preparation
- Hybridoma cell (>1×106 cells)
- Antigen (50 ug)
AbClon provided
- Variable region sequence
- Variable region DNA (H-L chain)
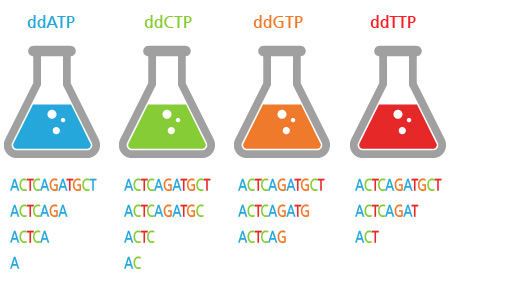
PCR process
- Overlap extension PCR
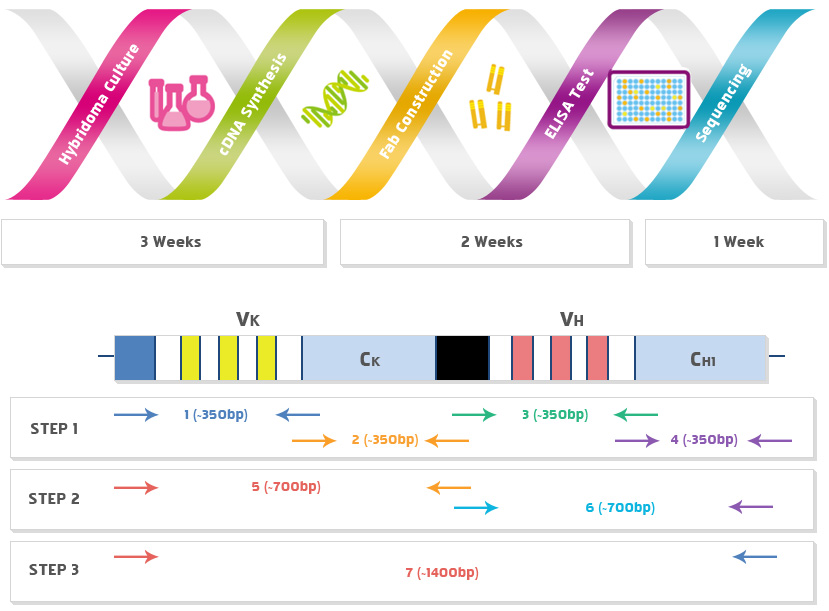
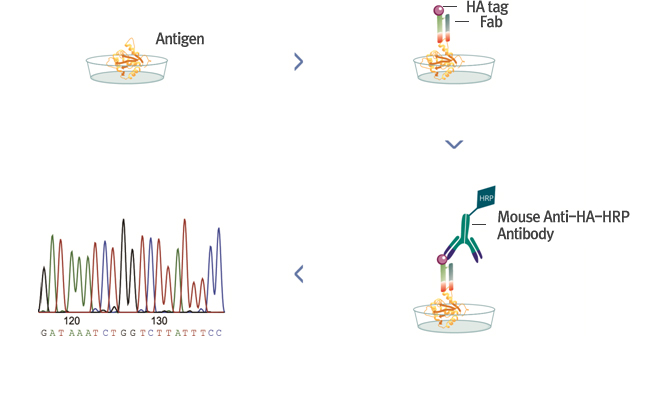
Identification after ELISA confirmation
- ∙ After confirming the reaction with the antigen by the Fab produced by PCR, the sequence is confirmed
Antibody humanization
Non-hHuman monoclonal antibody can beis more than 90% humanized while maintaining affinity and activity. The CDR grafting platform technology, which has know-how in the development of antibodies in AbClon, is accurate and reliable.
Features
- Reliable CDR grafting platform
- Duration: about 3 months
- Technology development for therapeutic antibody
Customer Preparation
- Variable region sequence
- The antigen (50 ug)
- IgG backbone (option)
AbClon provided
- Humanized sequence
- Variable region Humanized DNA (HL chain)
- Affinity result (KD)
- Full IgG (100 ug)
Service Flow
-
- Sequence analysis
-
- · Variable region
analysis
- · Variable region
-
- Frame work design
-
- · Frame work
comparison
- · Frame work
-
- CDR grafting
-
- · Human frame work
hzAb design
- · Human frame work
-
- Humanized
antibody production -
- · codon optimization
- · mammalian
expression
-
- Binding affinity
measurement -
- · octet
Affinity maturation
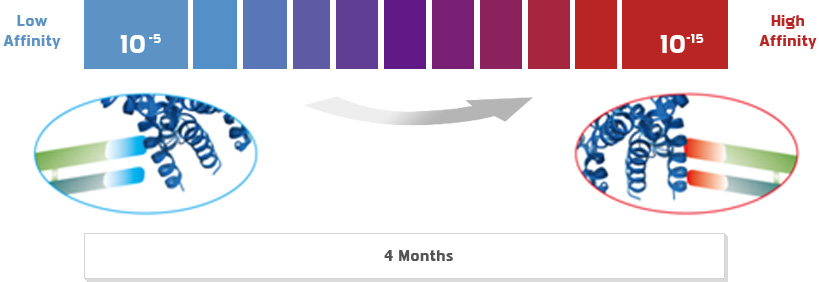
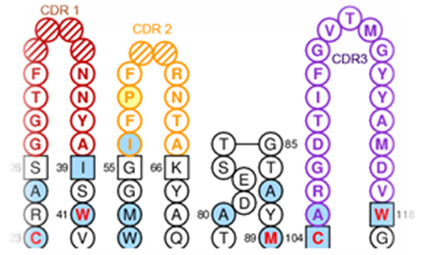
Naturally occurring affinity maturation in vivo occurs by somatic hypermutation and clonal selection. This principle and antibody library screening techniques are used to increase affinity in vitro. Biopanning improves affinity from libraries that have been granted diversity in the variable region. After measuring the affinity of the original antibody, the antibody library is developed by randomization for the CDR region in order to improve the affinity of the antibody, and the affinity-improved antibody is selected through the biopanning process. The diversity of the antibody library developed for affinity maturation is approximately 109 and affinity is analyzed using Octet.
Customer Preparation
- Variable region sequence
- The antigen (1 mg)
- Antibody (used as control)
- IgG backbone (option)
Offering an app clone (TOP 5 clones)
- Affinity matured sequence
- Affinity matured variable region DNA (HL chain)
- Affinity result (KD)
- scFv (100ug)
Service Flow
Building a Library
-
- CDR analysis
-
- · Check Affinity effective
-
- Library Design
-
- · Non-effective
randomization
- · Non-effective
-
- scFv library construction
-
- · Antibody library
-
- Verify library
-
- · Analysis of sequence
Affinity Selection
-
- Target binder and
sequence identification -
- · Panning
- · ELSIA
- · Sequencing
-
- Affinity measurement
-
- · Octet (scFv)
- · K-Off ranking TOP 5
-
- (scFv) production
-
- · 100ug
- · TOP 5
-
- KD Measurement
-
- · Octet (IgG)
+82-2-2109-1294
kdlee@abclon.com